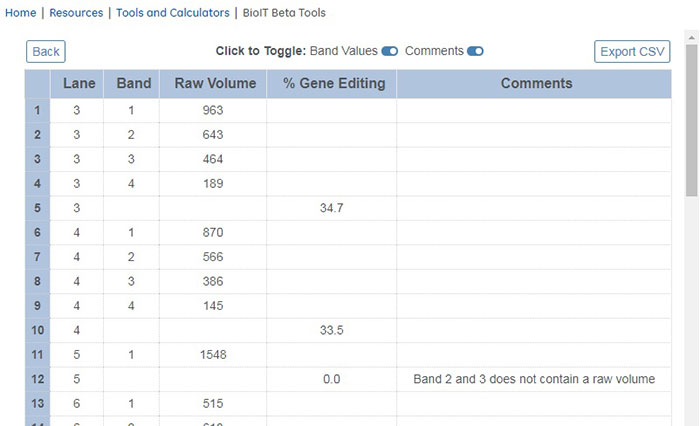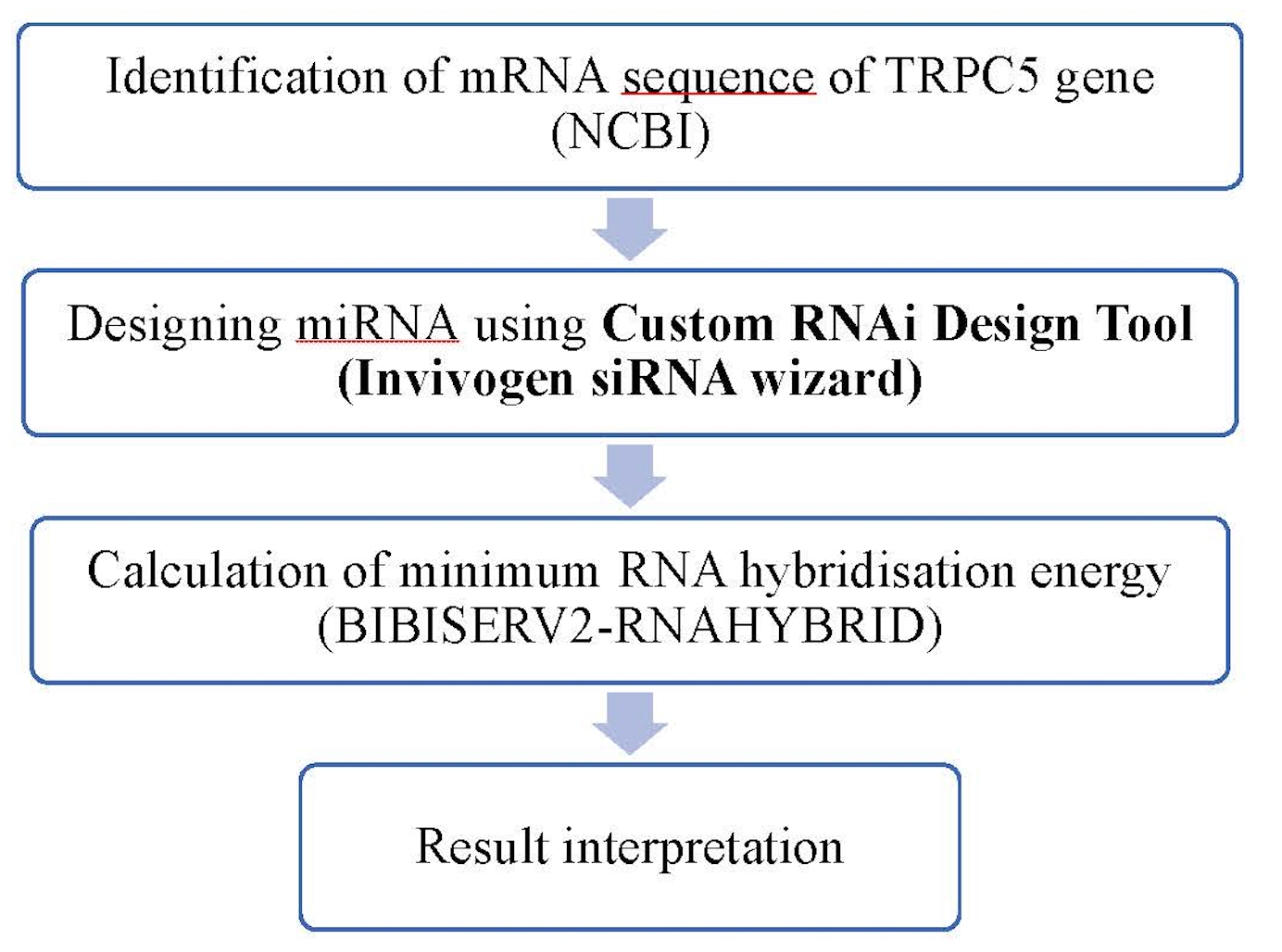
Assessment of siRNA as a therapeutic molecule in Transient Receptor Potential Channel 5 gene silencing: a computational approach | Biomedical Research and Therapy

Designing potential siRNA molecule for the nucleocapsid(N) gene silencing of different SARS-CoV-2 strains of Bangladesh: Computational approach - ScienceDirect

Molecules | Free Full-Text | A Comprehensive Computational Investigation into the Conserved Virulent Proteins of Shigella species Unveils Potential Small-Interfering RNA Candidates as a New Therapeutic Strategy against Shigellosis

CHARMM-GUI Free Energy Calculator for Practical Ligand Binding Free Energy Simulations with AMBER | Journal of Chemical Information and Modeling
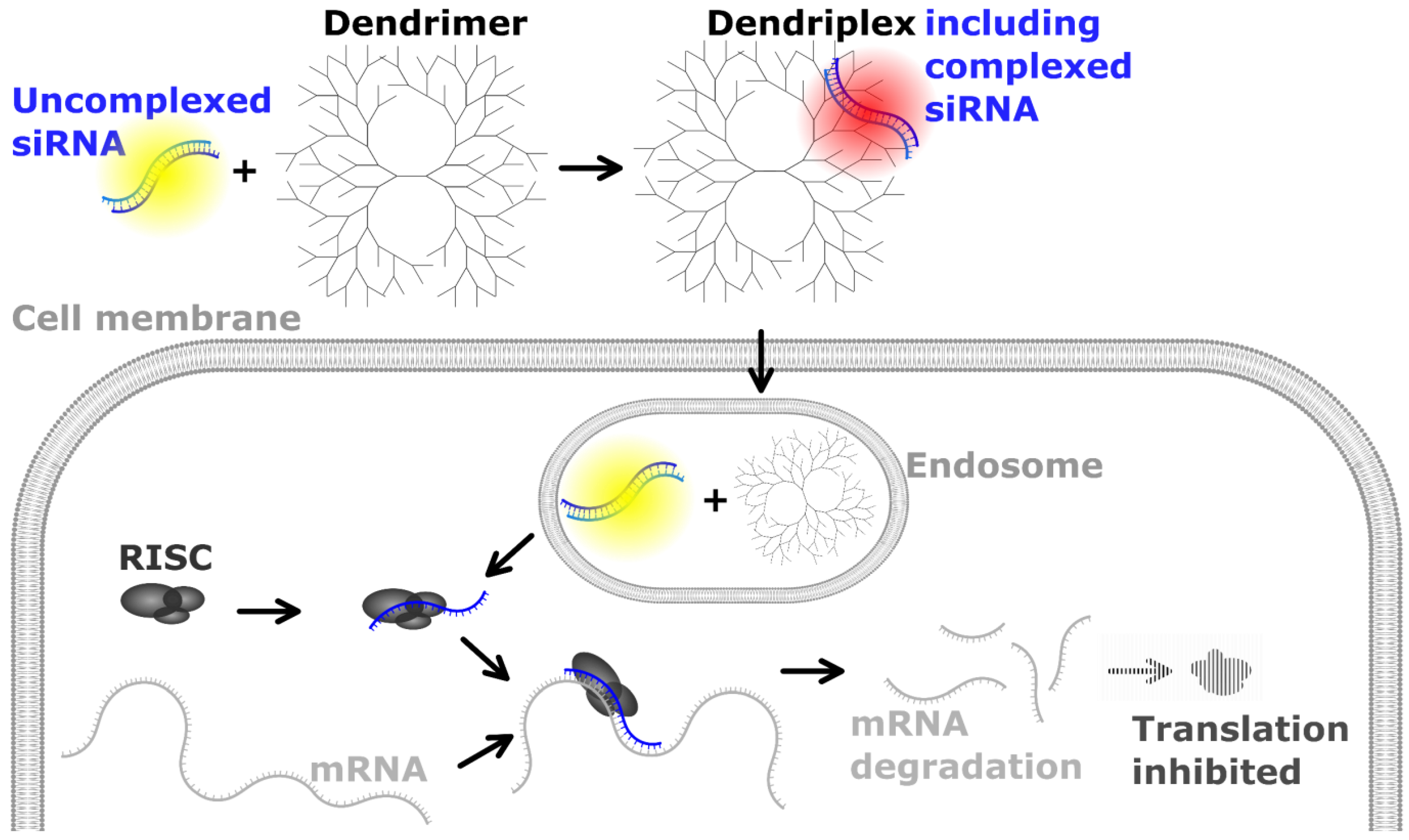
Pharmaceutics | Free Full-Text | Analyzing siRNA Concentration, Complexation and Stability in Cationic Dendriplexes by Stem-Loop Reverse Transcription-qPCR
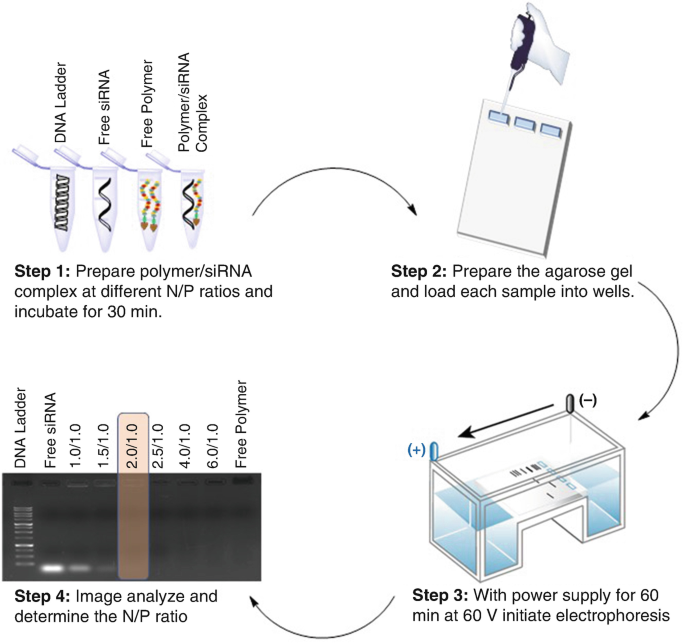
Determination of Optimum Ratio of Cationic Polymers and Small Interfering RNA with Agarose Gel Retardation Assay | SpringerLink

Morphology and size of PEI-D-GlcNAc-siRNA by TEM (A), size and size... | Download Scientific Diagram

CHARMM-GUI Free Energy Calculator for Practical Ligand Binding Free Energy Simulations with AMBER | Journal of Chemical Information and Modeling

Calculation of the probability of binding efficiency (PBE) based on... | Download Scientific Diagram

A Simple and Cost-Effective Approach for In Vitro Production of Sliced siRNAs as Potent Triggers for RNAi: Molecular Therapy - Nucleic Acids

Designing potential siRNA molecules for silencing the gene of the nucleocapsid protein of Nipah virus: A computational investigation - ScienceDirect
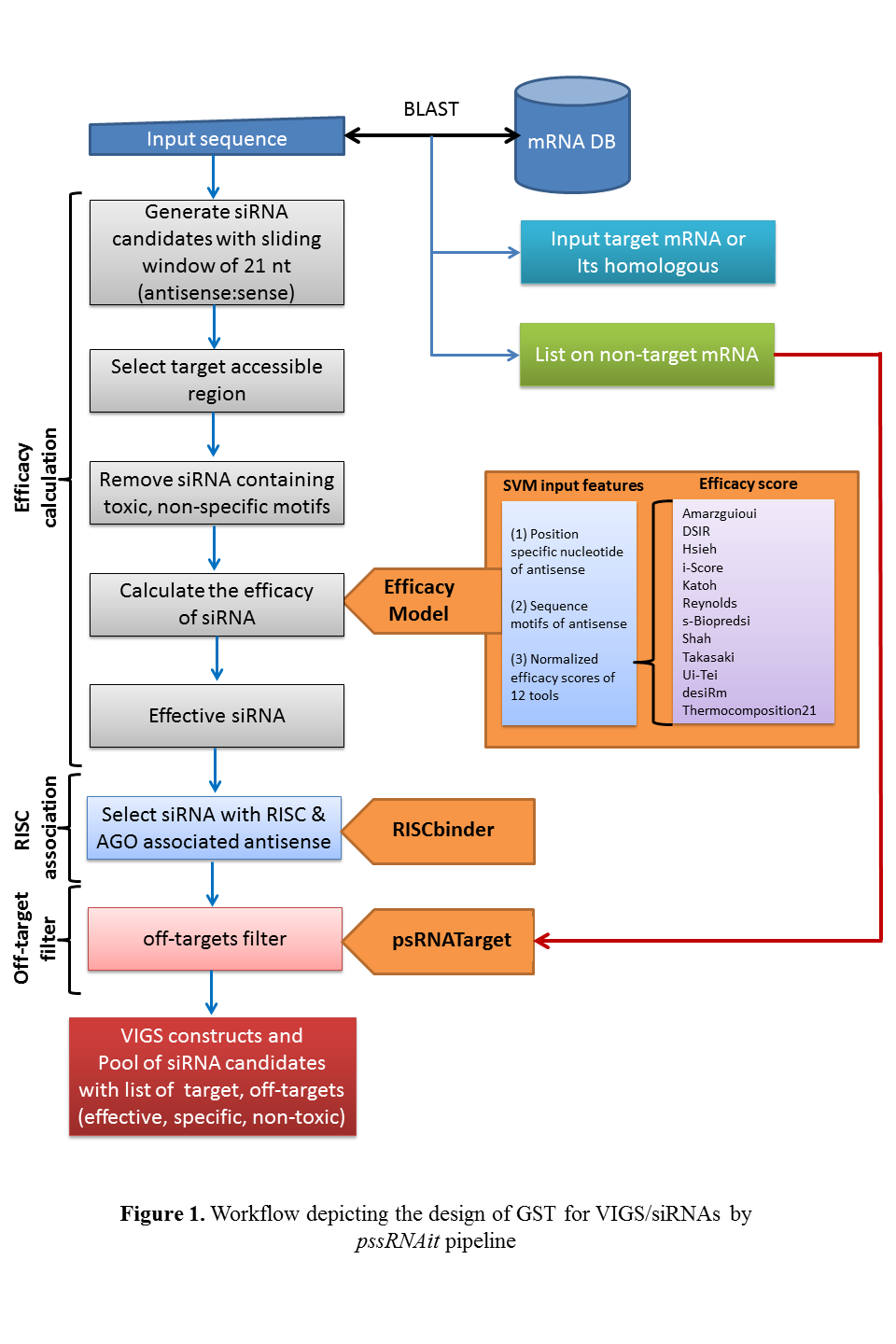
Welcome to pssRNAit: Designing Effective and Specific Plant RNAi siRNAs with Genome-wide Off-target Gene Assessment
![PDF] Demonstration of a ΔΔCq Calculation Method to Compute Relative Gene Expression from qPCR Data | Semantic Scholar PDF] Demonstration of a ΔΔCq Calculation Method to Compute Relative Gene Expression from qPCR Data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b42273036b0ed0ad4eba7b5012a1169d99e02b52/1-Figure1-1.png)
PDF] Demonstration of a ΔΔCq Calculation Method to Compute Relative Gene Expression from qPCR Data | Semantic Scholar